What are induced pluripotent stem cells (iPSCs)?
Induced pluripotent stem cells are adult cells, such as skin cells, that have been chemically reprogrammed in a lab to become a “master cell” with the ability to develop into any cell type in the human body. This reprogramming process allows scientists to create patient-specific stem cells to study diseases, model their progression, and test potential new therapies. This is where HeartWorks comes in!
Advancing Cardiac Therapies with iPSC Differentiation
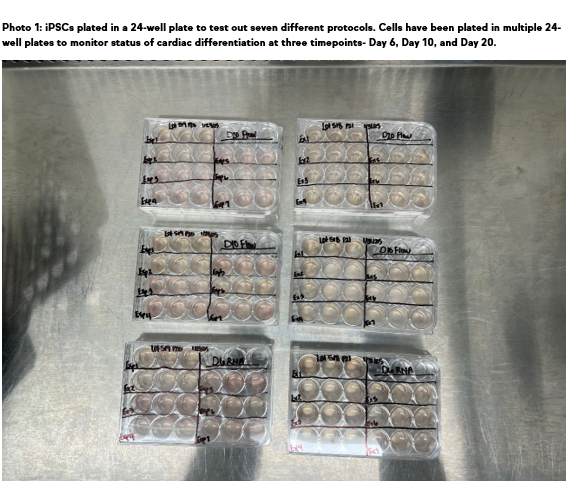
At HeartWorks, the Research and Development team has their focus on optimizing the production of cardiac-lineage cells from induced pluripotent stem cells (iPSCs). The 2D iPSC differentiation project is at the forefront of this effort, aiming to refine our protocols and ultimately enhance the efficiency and accessibility of our life-changing therapies.
Our current cardiac differentiation protocol sometimes falls short in consistently yielding the high percentage of cardiac-lineage cells we need. This can lead to multiple manufacturing attempts, increasing costs and, more importantly, potentially delaying critical treatments for patients. That’s where our 2D differentiation project comes in.
Why the 2D iPSC Differentiation Platform Matters
The 2D platform offers significant advantages for protocol optimization. By differentiating iPSCs in a flat, monolayer format, we can rapidly test numerous variations of our cardiac differentiation protocol. This high-throughput screening capability allows for quick cycles of learning, enabling us to pinpoint the precise conditions that lead to a higher percentage of healthy cardiac cells. This approach is also remarkably cost-efficient, requiring approximately 100 times fewer cells than experiments conducted in a 3D platform. The use of multi-well plates means our researchers can simultaneously execute multiple protocols with various replicates, accelerating our progress.
Since the project’s inception, we’ve already put 21 iPSC lines through 2D cardiac differentiation, analyzing over 115 samples from different protocol iterations. This extensive work is systematically guiding us toward an optimized protocol for clinical manufacturing.
From 2D to 3D: Translating Findings to Clinical Manufacturing
While the 2D platform offers incredible benefits for discovery, its primary challenge lies in its inherent difference from our 3D clinical manufacturing platform. In 2D, iPSCs differentiate as a monolayer, whereas in 3D, they form spheroids, which more closely mimic the body’s natural environment for cardiac development. The big question we face is: Can the promising results from our 2D-adapted cardiac-lineage cells truly translate to the clinically relevant 3D platform? Preliminary studies have yielded inconclusive results when testing the best 2D protocols in 3D culture, indicating a need for more in-depth understanding.
To address this, our team is embarking on a crucial time course study in the 2D platform. The goal is to meticulously examine gene expression patterns during the critical, early stages of cardiac differentiation in 2D. By comparing this data to existing time course data from our 3D platform, we aim to answer vital questions: Are there differences in the expression of key early cardiac genes? Does the timing of their expression vary? Are there different supporting or non-supporting cell types present? Unraveling these molecular differences is key to successfully translating our 2D findings to the 3D clinical setting.
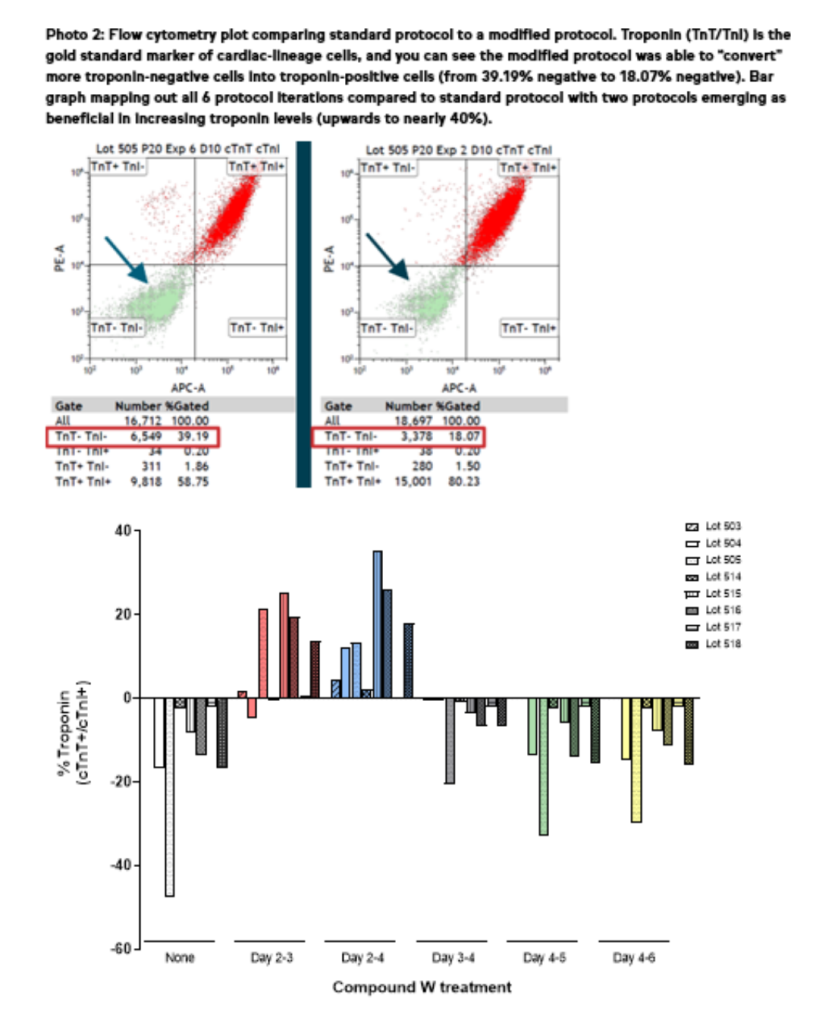
Despite the challenges, this project has already yielded exciting breakthroughs. We’re thrilled to report the emergence of two new cardiac differentiation protocols that have consistently outperformed our current standard, producing a higher percentage of cardiac-lineage cells. This significant achievement resulted from differentiating 3 iPSC lines, three times each, across seven different protocols that optimized the treatment time of a key signaling molecule.
The Future of Automated 2D Cardiac Differentiation at HeartWork
Looking ahead, we aim to maximize cardiac efficiency even more by combining these promising new protocols with more potent signaling molecules or those that target different pathways of cardiac differentiation. Another exciting prospect on the horizon is the development of an automated 2D cell culture platform. This could potentially revolutionize clinical manufacturing by allowing for the production of 2D-adapted cardiac-lineage cells, a significant pivot from our current 3D approach.
The breadth of knowledge gained and the possibilities emerging from the 2D differentiation project are truly rewarding. From developing improved protocols for clinical manufacturing to extensively characterizing gene expression pathways, it’s an incredibly exciting time to be part of discovery science here at HeartWorks. We are committed to continuing this vital work, ultimately bringing more efficient and accessible cardiac therapies to patients who need them most.
Email hello@webuildhearts.org with any questions!